Significance
Chemical biology tools have increasingly revealed the importance of sialic acids as a major signal in physiology and disease. α-2,6-Linked sialic acids on galactose drive cancer development and metastasis, immunological recognition, and microglial phagocytosis. This modification is biosynthesized by two enzymes: ST6-β-galactoside-α-2,6-sialyltranferase-1 (ST6GAL1), expressed throughout the human body, and ST6GAL2, predominantly seen in brain, breast, and colon. Although this modification is critical in both health and disease, the regulation and dysregulation of these enzymes and thus α-2,6-linked sialic acid are poorly understood. MicroRNAs can play a role in cancer development and are thought to exclusively suppress protein expression in dividing cells, such as tumor cells. But new research published in ACS Central Science shows that some of these tiny molecules can elevate the expression of a particular gene in dividing human cells and in cancer cells, challenging conventional wisdom.
One class of cellular machinery regulated by miRNAs are the enzymes involved in mediating glycosylation, which add carbohydrates to certain proteins. In cancer cells, however, this process can be highly dysregulated, suggesting that miRNAs could be doing something unusual. University of Alberta researchers led by Professor Lara Mahal set out to investigate exactly how miRNAs function within the glycosylation process, and whether the molecules might be functioning in a new way. The researchers developed a fluorescence assay that can analyze how miRNAs interact with their targets, and whether they increase or decrease the amount of protein produced. They used the assay to investigate the regulation of cancer-related glycosylation enzymes ST6GAL1 and ST6GAL2, and found that for the former, the miRNAs appeared to directly upregulate the process in noncancerous human cells.
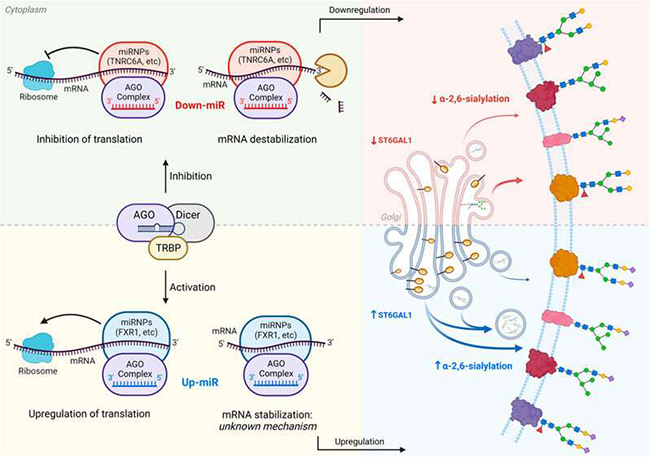
Glycosylation is the outcome of a complex regulatory network in which biosynthetic enzymes can be modulated at the transcriptional, post-transcriptional, translational, and post-translational levels. miRNAs are important modifiers of protein expression, engaging with the 3′-untranslated region (3′-UTR) of mRNA to tune expression levels. The canonical view of this interaction is that it represses protein levels through the formation of RISC complexes comprised of Argonautes (AGOs), a variety of RNA binding proteins, mRNA and miRNA. The exact nature and number of these complexes are unknown. Repression can occur through either destabilization of the message or inhibition of translation itself. In nondividing cells and in mitochondria, select cases of miRNA activation of protein expression via direct interactions with the 3′-UTR have also been observed. This upregulation is thought to require destabilized mRNA lacking a 5′-cap and a typical poly(A) tail. These conditions are not met by typical mRNA in actively dividing cells, such as cancer cells, and upregulation of protein expression by miRNA is not thought to occur under these circumstances. Here we show that, in contrast to current assumptions, miRNA can upregulate protein expression, and corresponding glycosylation, in proliferating cancer cells.
The research team recently established a high-throughput fluorescence assay, miRFluR, that enables interrogation of the entire cohort of human miRNA against the 3′-UTR of a single gene. The authors comprehensive analysis showed intriguing hints that upregulation of protein expression might not be limited to nondividing cells. They applied the new assay to analyze the regulation of ST6GAL1 and ST6GAL2, the enzymes underlying α-2,6-sialic acid. Their findings revealed a surprising result, namely, that most miRNAs impacting ST6GAL1 upregulate the enzyme and α-2,6-sialylation in proliferating cells. Upregulatory miRNAs were also observed for ST6GAL2, although in this case it was not their major mode of action. The authors showed that upregulation by miRNA occurs through direct binding between the miRNA and 3′-UTR. The new study may help explain why α-2,6-sialylation is commonly upregulated in cancer, as miRNAs that upregulate ST6GAL1 are high in cancers, such as pancreatic cancer, that have high levels of α-2,6-sialylation. Overall, the new study showcases the power of chemical biology to bring new understanding to fundamental questions.
In many cancers, α-2,6-linked sialic acids are overexpressed, and dysregulation of this glycan is emerging as a crucial part of cancer formation, metastasis, and immune recognition. miRNA are major regulators of the glycome, but their role in controlling α-2,6-linked sialic acid has not been well-studied. The analysis of the miRNA regulatory landscape for the α-2,6-linked sialylation enzymes ST6GAL1 and ST6GAL2, described herein, has revealed new potential links between miRNA and the upregulation of α-2,6-linked sialosides observed in cancer.
The dominant view of miRNA regulation is that in proliferating cells the direct impact of miRNA on protein expression is downregulatory. The high-throughput analysis of ST6GAL1 and ST6GAL2 contradicts this, revealing that upregulatory interactions may be commonplace. Consistent with this, a smaller high-throughput luciferase assay for POT1, PTEN, MXI1, and other cancer-related genes also identified a number of upregulatory miRNA interactions, but these were ignored as noise. Previous miRFluR analysis of miRNA-mediated regulation for B3GLCT also identified potential up-miRs for that enzyme. These interactions have been missed by the scientific community because the current pathway for identifying miRNA interactions has depended on validating potential targets of miRNAs predicted by Targetscan and other algorithms that are focused on downregulation. Recent work knocking out AGO complexes found that removal of the miRNA machinery caused most genes to lose expression, consistent with upregulation being a primary function of miRNA, rather than the expected gain that would come from loss of a repressor. Taken together, the data support upregulation as part of the broader landscape of miRNA regulatory mechanisms in both dividing and quiescent cells.
For ST6GAL1, which is known to be upregulated in many cancers, upregulation appears to be the major mode of action of miRNA, although these same miRNAs have downregulatory activity for other genes. Given the importance of miRNA in tuning the expression of genes, it is perhaps unsurprising that regulation by miRNA would be in both directions. Precise control over protein expression is critical for low abundance proteins, where noise becomes an increasing problem. This class of proteins includes many glycosylation enzymes, GPCRs, and most cell surface receptors. These proteins, which often act as initiators of amplified signals, would be important to tightly regulate. The expanded understanding of the miRNA regulatory landscape opens new possibilities for miRNA mechanisms to modulate protein expression and create tools to further explore the impact of these noncoding RNA. Furthermore, this challenges the current understanding that miRNAs only downregulate protein production. They also tested for miRNA-mediated upregulation in multiple cancer cell lines and observed the same results. The researchers say that this work expands the understanding of how miRNAs work, an important consideration for using miRNA-based therapeutics in both current and future clinical trials.
Reference
Faezeh Jame-Chenarboo, Hoi Hei Ng, Dawn Macdonald, and Lara K. Mahal. High-Throughput Analysis Reveals miRNA Upregulating α-2,6-Sialic Acid through Direct miRNA–mRNA Interactions. ACS Central Science (2022). DOI: 10.1021/acscentsci.2c00748
 Medicine Innovates Medicine Innovates: Delivering innovations in medicine to the world for better health and prosperity
Medicine Innovates Medicine Innovates: Delivering innovations in medicine to the world for better health and prosperity

