Significance
Acute Myeloid Leukemia (AML) is marked by the rapid proliferation of abnormal hematopoietic stem cells, leading to a high relapse rate despite initial treatment success. Traditional treatment has involved anthracyclines and cytosine arabinoside (AraC), known collectively as conventional chemotherapy. More recently, targeted therapies like the combination of Venetoclax with Hypomethylating agents (VH) have shown promise. Despite these advancements, the heterogeneity in patient response necessitates tailored therapeutic strategies. To this end, a new study published in Leukemia and conducted by Dr. Eduardo Sabino de Camargo Magalhães, Dr. Stefan Edward Hubner, Dr. Brandon Douglas Brown, Dr. Yihua Qiu & led by Professor Steven Mitchell Kornblau at the University of Texas MD Anderson Cancer Center, the researchers ran a proteomic-based approach using Reverse-phase Protein Arrays (RPPA) for enhancing therapy selection between VH and conventional chemotherapy in AML patients. They aimed to refine treatment selection between VH and conventional chemotherapy by employing a proteomic profiling strategy to identify predictive protein signatures. The analysis of 810 patients revealed distinct protein profiles that could potentially guide treatment decisions, increasing overall survival rates and the possibility of achieving remission.
First, the researchers collected blood and bone marrow samples from newly diagnosed adult AML patients at the University of Texas MD Anderson Cancer Center. Samples were processed shortly after collection, using Ficoll gradient centrifugation to separate mononuclear cells, followed by Magnetic Activated Cell Sorting (MACS) to deplete T and B cells. Afterward, samples were lysed to extract proteins, which were then denatured and arrayed on nitrocellulose-coated slides in a series of five 2-fold serial dilutions. Slides were probed with 411 validated antibodies (including both total and phosphorylated proteins) and detected using secondary antibodies linked to an infrared dye. The stained slides were scanned to quantify protein levels, with data normalized against protein levels in normal bone marrow-derived CD34+ cells. The team employed machine learning algorithms (primarily random forests) to analyze the protein expression data. The aim was to develop a classifier capable of predicting which treatment modality (VH or conventional chemotherapy) would be most effective for each patient. Using expression data, they identified 109 proteins with significant prognostic value. Further analysis refined this to 14 key proteins that effectively discriminated between optimal treatment regimens. The model’s predictions were validated against clinical outcomes, focusing on overall survival and treatment efficacy.
The team successfully identified 109 proteins that had prognostic value in AML. These proteins were used to categorize patients into five distinct expression profiles that correlate with different clinical outcomes. A subset of 14 out of the 109 proteins was found to be particularly effective in predicting the best treatment approach. This classifier can be used to guide the choice between VH and conventional chemotherapy, aiming to maximize patient response and survival. Implementing the proteomic classifier was projected to lead to a change in therapy for about 30% of the patients. This adjustment was estimated to increase the 5-year overall survival rate by 43%, translating to approximately 2600 additional cures annually in the United States. Beyond the immediate application in AML, this approach has potential implications for other types of cancer where treatment optimization remains a challenge. The method’s scalability and the detailed molecular insights it provides could facilitate wider adoption in personalized medicine. Importantly, the study also identified a group of patients who did not benefit from either VH or conventional chemotherapy, suggesting the need for alternative therapeutic strategies for this subgroup.
The significance of the study lies in its potential to revolutionize the treatment of AML through the use of proteomic profiling and machine learning algorithms. First, the study proposes a method to tailor AML treatment plans based on individual protein expression profiles, moving away from a one-size-fits-all approach and towards more personalized medicine. This has the potential to significantly increase the effectiveness of treatments by matching patients with the therapies most likely to benefit them. Secondly, by identifying specific protein signatures that correlate with better responses to either Venetoclax combined with Hypomethylating agents or conventional chemotherapy, the study suggests that it could be possible to increase overall survival rates and complete remission durations for AML patients. This would represent a major advance in a disease area where outcomes have historically been poor. Thirdly, by more accurately predicting which patients will benefit from which treatment regimens, unnecessary exposure to potentially ineffective and toxic treatments can be minimized. This not only improves quality of life but also reduces healthcare costs associated with ineffective treatment. Moreover, the study enhances understanding of the molecular underpinnings of AML through detailed proteomic analysis. Such insights are invaluable for the development of new therapeutic targets and improving existing treatments. Furthermore, using RPPA technology combined with machine learning offers a scalable method for analyzing protein expression in other cancers and diseases, suggesting that this approach could have broad applications beyond AML. The study estimates that implementing its findings could lead to around 2600 more cures annually in the United States alone, underlining its significant potential impact on public health.
Overall, the integration of RPPA-based proteomic profiling into clinical practice could significantly impact the management of AML by providing a more nuanced understanding of disease biology and patient-specific therapeutic responses. The new method reported by Professor Steven Mitchell Kornblau and his team holds promise for improving patient outcomes through tailored treatment strategies, thus marking a significant advance in the personalized treatment of AML.
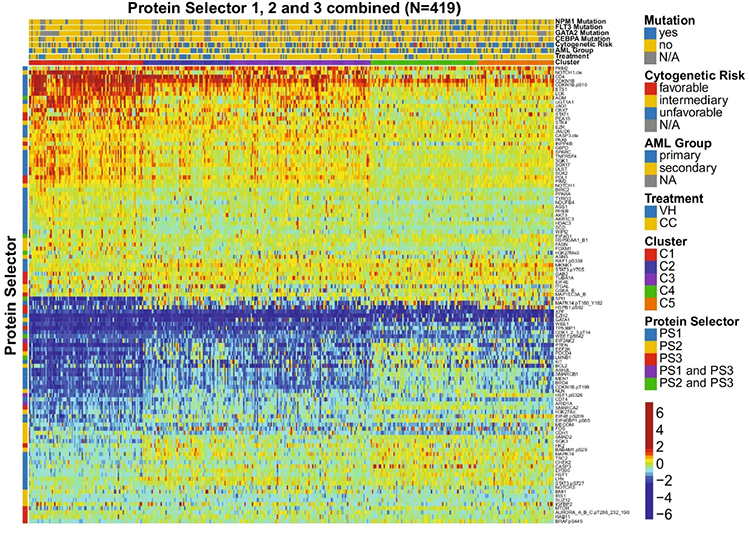
Reference
de Camargo Magalhães ES, Hubner SE, Brown BD, Qiu Y, Kornblau SM. Proteomics for optimizing therapy in acute myeloid leukemia: venetoclax plus hypomethylating agents versus conventional chemotherapy. Leukemia. 2024 Mar 26. doi: 10.1038/s41375-024-02208-8.
 Medicine Innovates Medicine Innovates: Delivering innovations in medicine to the world for better health and prosperity
Medicine Innovates Medicine Innovates: Delivering innovations in medicine to the world for better health and prosperity

